If you want something that you can script in order to generate some nice renderings I can also suggest PyMOL and VMD. IQmol can also directly open some Gaussian files to visualize frequencies, however, since it's been just recent that I obtained Gaussian on my PC I don't have that much experience here, yet. step plot) using the *_trj.xyz file that is automatically generated during an optimization run in Orca and you can visualize frequencies after generating the xyz file using the orca_pltvib tool. This way you can monitor the geometry optimization (including an energy vs. If you have xyz files containing multiple geometries they can be visualized as animations. 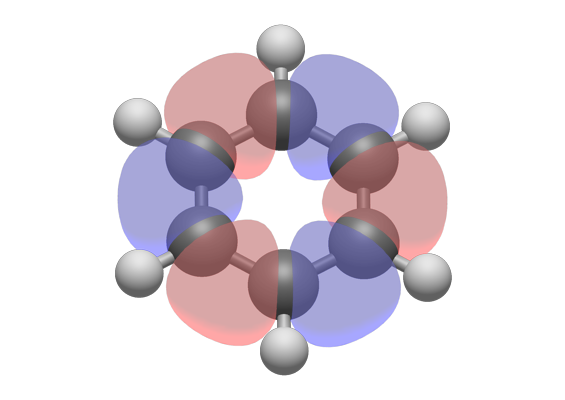
and it's super easy to make high quality renderings for reports etc. Since I've been using Orca mostly I've been using the orca_plot tool to generate cube files that can be opened in IQmol in order to visualize densities, orbitals, etc. Solution: A The molecular mass of ethylene glycol can be calculated from its molecular formula using the method illustrated in Example 3.2.1: 2C(2atoms)(12.011amu / atom) 24.022amu. It runs perfectly and has multiple force fields implemented for generating initial geometries. Convert from moles to molecules by multiplying the number of moles by Avogadro’s number. Since for any substance there are 6. I ended up using IQmol as an alternative. This conversion factor involves the number of particles per unit amount of substance and is given the symbol NA N A and called the Avogadro constant. It's been a while since I've tried it the last time so I would have to check what the problem was and if it still exists. For example, the molecular weight of oxygen is 32.00, so that one gram-mole of oxygen has a mass of 32.00 grams and contains 6.02214076 × 10 23. Yes, I experienced very similar problems. The specific number of molecules in one gram-mole of a substance, defined as the molecular weight in grams, is 6.02214076 × 10 23, a quantity called Avogadro’s number, or the Avogadro constant. Is this a a problem on my end, or are there some issues with Avogadro2 in arch linux? Has any of you faced these issues and eventually managed to get it to work smoothly? Has any of you managed to install the original Avogadro in arch? Or, is there any other FOSS molecular viewer that you recommend for building small molecules and quickly optimizing the geometries using forcefield methods, visualizing output MOs, and frequencies in Arch Linux? I have found that Avogadro2 does not do these things as smoothly, but most importantly, I find that it is very laggy and it crashes often. I have used Avogadro in the past for quickly building starting geometries, visualizing the output geometries, frequencies, and orbitals. 
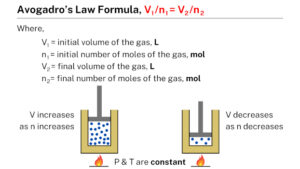
Where, The pressure exerted by that of the gas on the walls from its container is denoted by ‘P,’ and the volume occupied by gas is denoted by ‘V.’. The ideal gas equation, that can be articulated as follows, can be used to derive Avogadro’s law: PV nRT. In Avogadro 1.9x it’s under Build > Atomic Coordinate Editor. Avogadro’s Equation from Ideal Gas Equation. It would be nice to have back the Avogadro. Below is general information regarding the force fields to help you select the best optimization method. Avogadro comes equipped with multiple different force fields. AMBER (Assisted Model Building with Energy Refinement) is a common protein force field. Well there is quite a bit of information for the modes. GAFF (General AMBER Force Field) is often used for optimizing the geometries of drugs. right now if we compute with NumFreq option and look for Raman and or IR the output looks like shown below. In Avogadro 1.2, it’s under Build > Cartesian Editor. Avogadros number is defined as the number of elementary particles (molecules, atoms, compounds, etc.) per mole of a substance. I have tested the ORCA 5.0.3 output vs Avogadro 1.97, it does not seem to work, might be a change of formating which Avogadro looks for. You can paste this into the Cartesian Editor as well. For instance, consider the size of one single grain of wheat. It is quite difficult to visualize a mole of something because Avogadro's constant is extremely large. is analogous to saying: 1 Dozen 12 eggs (2) (2) 1 Dozen 12 eggs. Eventually I settled for installing Avogador2. I also tried copying just the coordinates of the final geometry. Using the following relation: 1 mole 6.02214179 ×1023 (1) (1) 1 mole 6.02214179 × 10 23. I have changed my OS to Arch Linux, and I was not able to build the older avogadro because of multiple compatibility issues. In the SI system of units, the value of the elementary charge is exactly defined as e īy combining the best measured value of the antiproton charge (below) with the low limit placed on antihydrogen's net charge by the ALPHA Collaboration at CERN.I used to work in a Ubuntu PC using the original version of Avogadro in Ubuntu. The elementary charge, usually denoted by e, is a fundamental physical constant, defined as the electric charge carried by a single proton or, equivalently, the magnitude of the negative electric charge carried by a single electron, which has charge −1 e.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
